The SAS team is interested in specialized metabolites synthesized by plants that have a hormonal function and/or are involved in interactions between plants in the rhizosphere (allelopathy). The team aims to identify these metabolites and decipher their synthesis and signalling pathways.
Biological Question
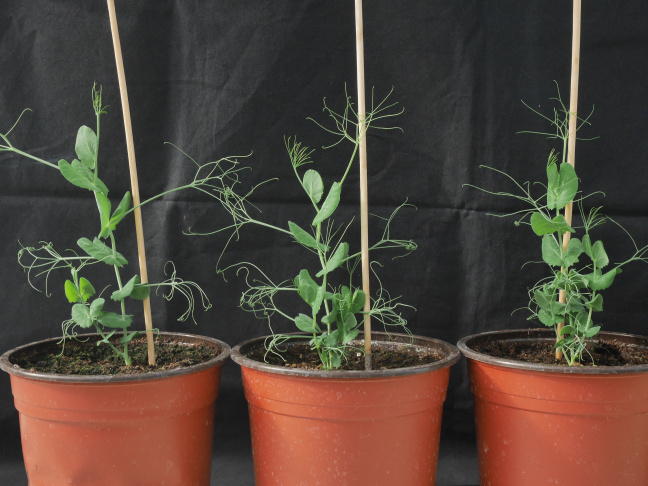
Plants synthesize an immense diversity of specialised metabolites (formerly known as secondary metabolites) that may have functions in development and growth, defence or ecological functions (interactions with soil micro-organisms). The SAS team has been working for many years on strigolactones (SL), the 9th class of hormones identified in 2008 for their role in controlling branching. These molecules, derived from carotenoids, were already known for their role in the rhizosphere. They act as germination stimulants for the seeds of parasitic plants such as Orobanche and Striga. SLs also play a key role in establishing symbiosis with arbuscular endomycorrhizal fungi. This ancient symbiosis, which is observed in over 80% of extant land plants, is postulated to have played a pivotal role in the colonization of the terrestrial environment by plants over 450 million years ago.
Models, tools and methods
Societal and economical impacts
Plants synthesize an immense diversity of specialised metabolites (formerly known as secondary metabolites) that may have functions in development and growth, defence or ecological functions (interactions with soil micro-organisms). The SAS team has been working for many years on strigolactones (SL), the 9th class of hormones identified in 2008 for their role in controlling branching. These molecules, derived from carotenoids, were already known for their role in the rhizosphere. They act as germination stimulants for the seeds of parasitic plants such as Orobanche and Striga. SLs also play a key role in establishing symbiosis with arbuscular endomycorrhizal fungi. This ancient symbiosis, which is observed in over 80% of extant land plants, is postulated to have played a pivotal role in the colonization of the terrestrial environment by plants over 450 million years ago.
Our team aims to study the functions and signalling pathways of SL during evolution and seeks to answer the following questions: How did the different functions of SLs emerge over the course of evolution? How have SL receptors and signalling pathways evolved between non-vascular and vascular plants? Which ligand(s) is (are) perceived by the ancestor of the SL receptor, the KAI2 protein?
More recently, the team has embarked on a project to identify and characterize other rhizosphere metabolites involved in plant-plant interactions, a phenomenon known as allelopathy. Specialized metabolites (formerly called secondary metabolites), when produced at the root level and released into the rhizosphere, can have very marked effects (positive and negative) on neighboring plants or soil microorganisms. The team is aiming to identify new metabolites and associated genes involved in alleopathic processes.
Models, tools and methods
Our current projects focus on the mechanisms of SL perception and signalling in vascular plants (Arabidopsis, pea, and parasitic plants such as Orobanche cumana), and in a bryophyte, the moss Physcomitrium patens. We are using complementary approaches in genetics, physiology, cell biology and biochemistry. Our collaboration with a chemist from the ICSN (François-Didier Boyer, CNRS Gif sur Yvette) has been very active for many years and remains at the heart of our projects.
The more recent project on allelopathy uses the many advantages of the model species Arabidopsis and the genetic resources available to identify the genes involved in interactions (positive and negative) between plants using association genetics approaches. The metabolites involved in these interactions are sought by non-targeted analysis of root exudates from Arabidopsis accessions and mutants.
Societal and economical impacts
The development of innovative strategies for tomorrow's agriculture, through the selection of plants adapted to agroecology, is based on knowledge of plants' natural defences and the molecular mechanisms that regulate their interaction with their environment. Our projects are fully in line with the current context of diversification of agro-systems and the gradual reduction in the use of pesticides and herbicides: - to combat parasitic plants of the Orobanchacea family, such as Striga and Orobanche, which are devastating in Africa, in Mediterranean countries and are beginning to cause major problems in France for oilseed rape (Phelipanche ramosa) and sunflower (Orobanche cumana); - to select varieties that suppress the development of weeds or genotypes adapted to plant associations.

Leader:
Sandrine Bonhomme