The team SYNAPS dedicates its research on finding solutions that favor the agroecological transition. More specifically, the team’s major objective is to understand the molecular and physiological mechanisms improving crop Nitrogen Use Efficiency (NUE) mediated by beneficial microorganisms.
Welcome to the SYNAPS team webpage!

Scientific goals :
The SYNAPS team focuses its research activities on developing solutions that support the agroecological transition. Especially, our primary objective is to investigate the molecular and physiological mechanisms by which beneficial microorganisms enhance crop Nitrogen Use Efficiency (NUE) (Figure 1).
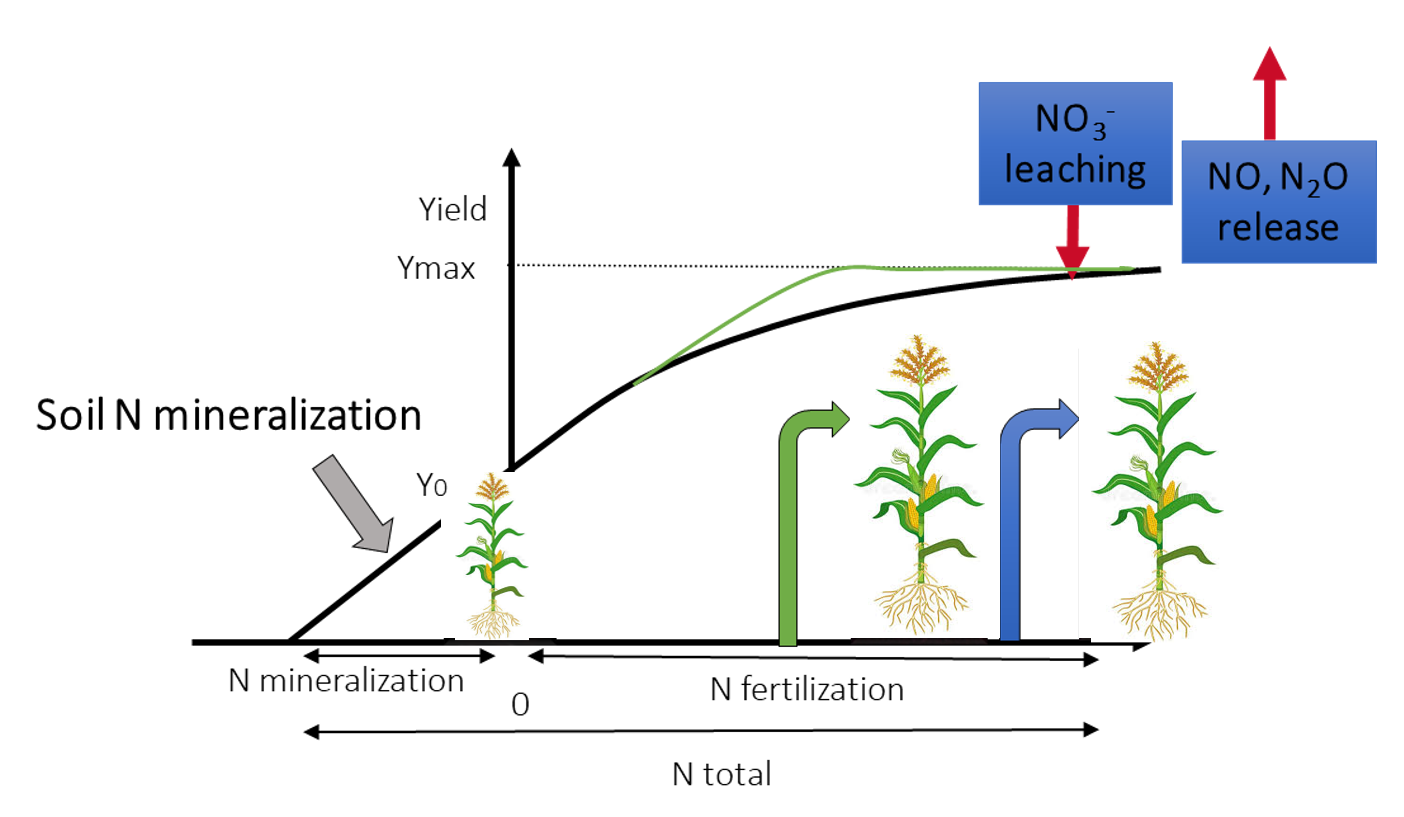
Figure 1: Nitrogen Use Efficiency (NUE) allows determining the capacity of a crop to manage the available N to produce grain yield or biomass yield. The plant with the green arrow has a better NUE than the one with the blue arrow. Higher NUE leads to less fertilizer use and less environmental pollution. Several levers can improve crop NUE including breeding, crop rotations, intercropping and exploiting microbes to increase N availability.
The team implements system biology approaches, using model crops, integrated multi-omics, genomics, metabolic modelling and microbial ecology to characterise plant N nutrition mediated by its microbiome. Our research is focused on the microbiome-mediated N nutrition of cereals such as maize and sorghum alone or intercropped with legumes such as soybean (Figure 2) .
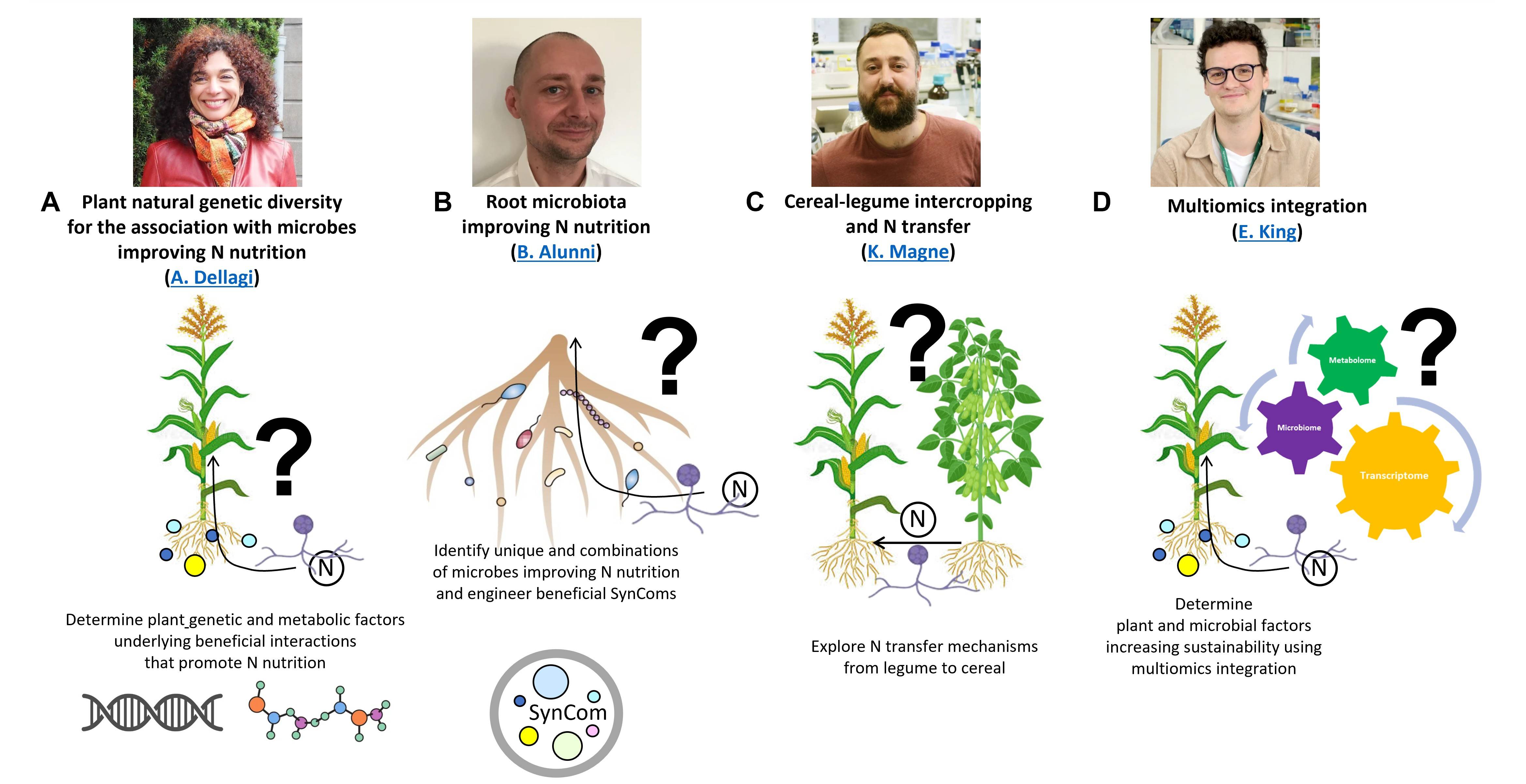
Figure 2: Four main axes of research for a sustainable improvement of N use. A) Understanding the genetic, physiological and metabolic mechanisms taking place in the plant and in the microbes that improve N use (Alia DELLAGI). B) Exploring and engineering crop microbiome by creating microbial Synthetic Communities (SynComs) based on their metabolic complementarities in boosting AMF mediated N crop nutrition (Benoît ALUNNI). C) Understanding genetic, physiological and metabolic mechanisms taking place in cereal-legume intercropping systems and the role of the microbiome in favoring N transfers via the common mycorrhizal network (Kévin MAGNE). D) D) Integrate multi-omics to identify microbial and/or plant factors for a sustainable production (Eoghan KING).
Our biological questions:-
A) What are the physiological, metabolic and molecular mechanisms involved in maize N nutrition during symbiosis with AMF? We demonstrated that maize interaction with the arbuscular mycorrhizal fungus (AMF) Rhizophagus irregularis alleviates maize N deficiency stress (Figure 3) and are developing a series of biological and experimental tools to understand by which mechanisms this process takes place. We carry out and integrate multi-omics approaches, including transcriptomic, metabolomic, fluxomic and ionomics data at the level of plant organs and aim at including the microbial compartments. Using microscopy we track the fate of arbuscules during symbiosis (Figure 4) trying to determine whether their decay may provide N to the plant. These approaches will help to better understand the physiological and molecular processes underlying maize-AMF interactions which improve maize NUE.
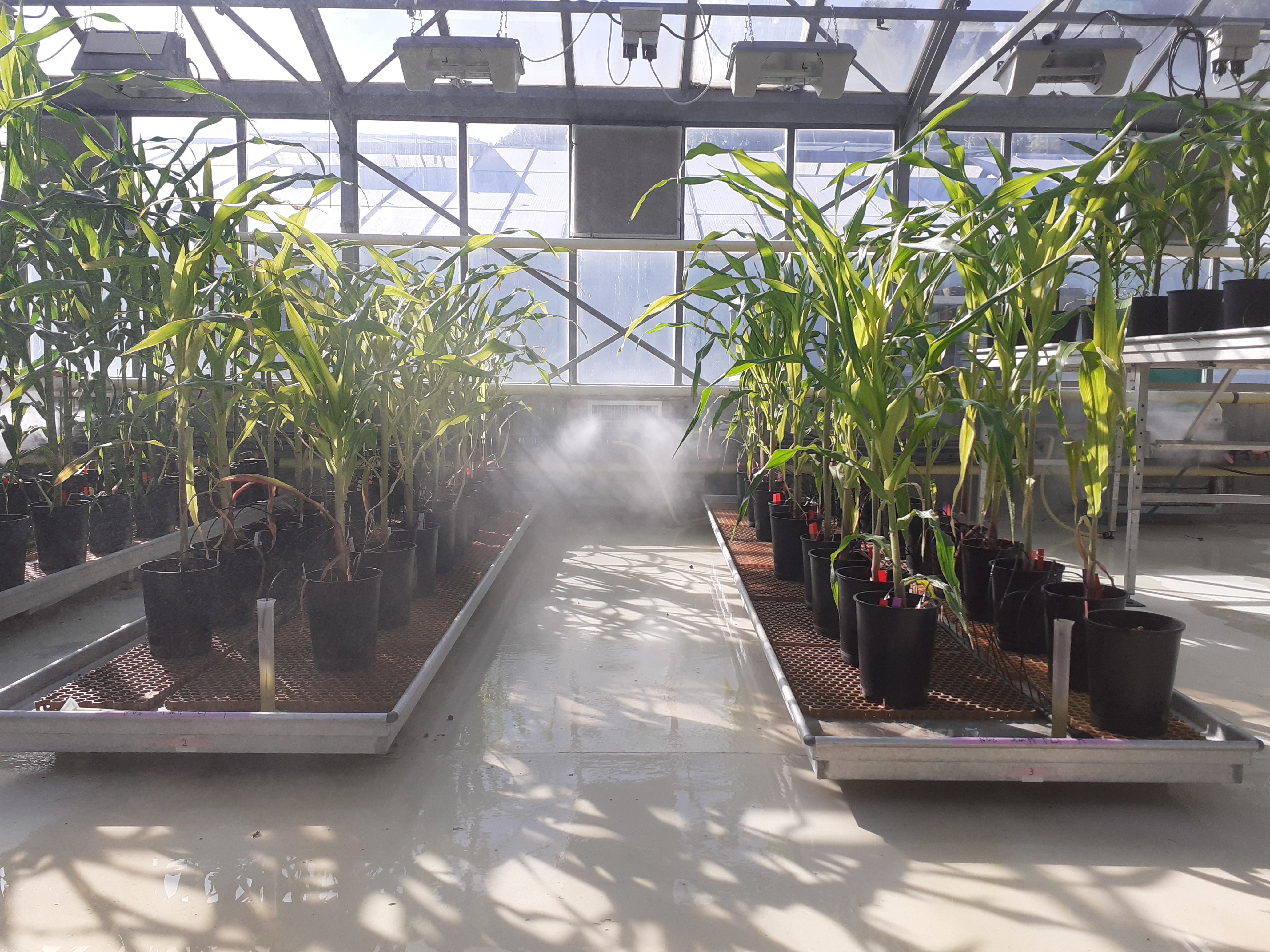 Figure 3: Maize plants grown under limiting N conditions and inoculated with the AMF R. irregularis (right) or not (left).
Figure 3: Maize plants grown under limiting N conditions and inoculated with the AMF R. irregularis (right) or not (left).We implemented system biology approaches based on multi-omics analysis and metabolic modeling tools. Owing to a collaboration with Dr Rajib SAHA (Univ. Nebraska) our study supports the idea that the pyrimidine metabolic pathway is a crucial component of maize N metabolism during AM symbiosis under N-limited conditions (Decouard et al., 2023, BioRxiv, Chowdhury et al. 2023). In collaboration with the IJPB team NUTS, we are investigating the response of Brachypodium distachyon to R. irregularis under low N condition. In collaboration with the IJPB team QALIBIOSEC we are investigating the impact of plant cell wall composition on AM symbiosis and vice versa.
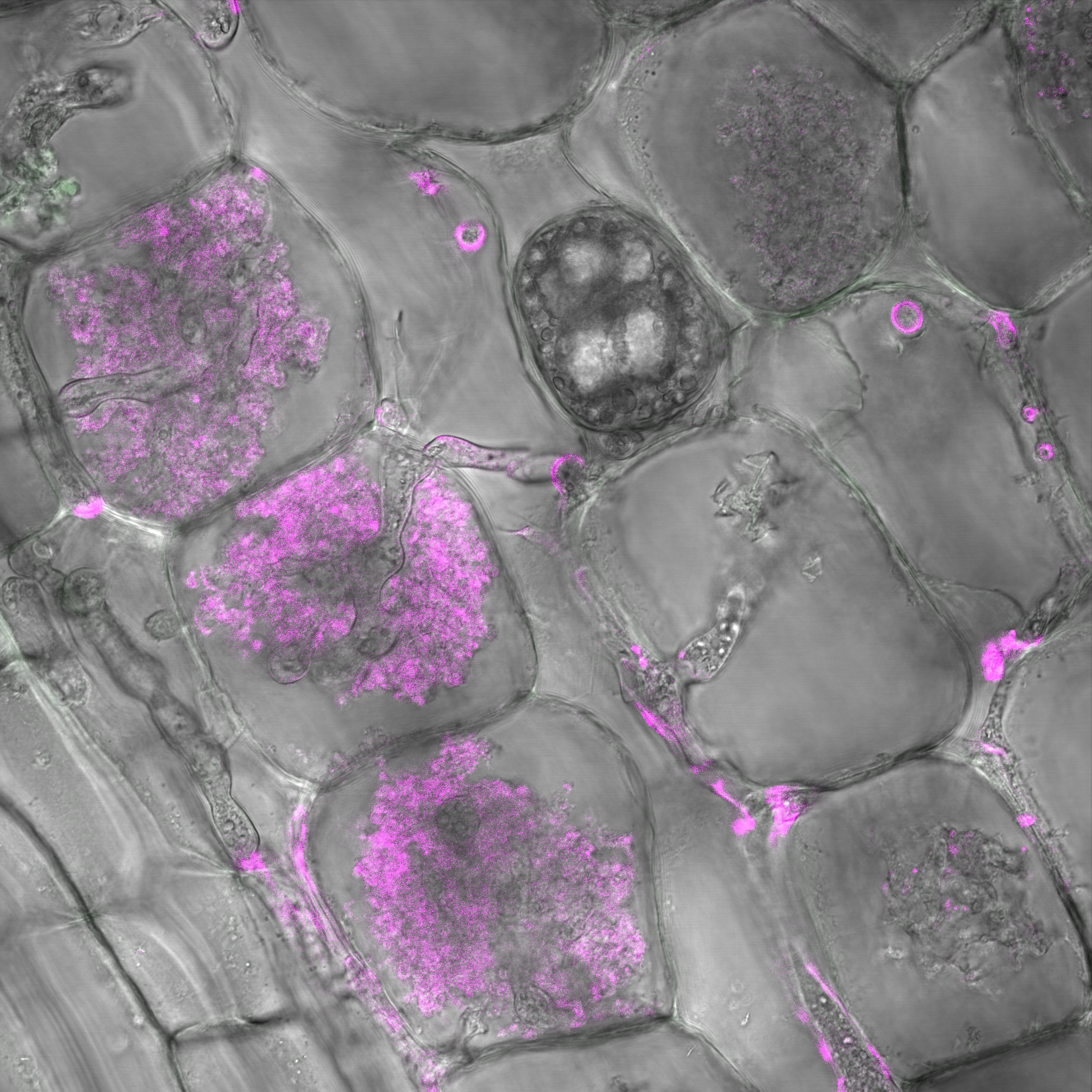 Figure 4: W22 maize roots colonized by arbuscules. R. irregularis arbuscules are highlighted using Wheat Germ Agglutinin (WGA)-Alexa Fluor 647. Magenta structures are AMF arbuscules and hyphae.
Figure 4: W22 maize roots colonized by arbuscules. R. irregularis arbuscules are highlighted using Wheat Germ Agglutinin (WGA)-Alexa Fluor 647. Magenta structures are AMF arbuscules and hyphae.B) What are the microbial consortia that can boost maize N nutrition by associating AMF and by which mechanism ? This research will delve into mechanisms of maize microbiota recruitment by root exudates and the characterization of the microbial consortia associated with AMF-associated maize root boosting N nutrition. To address these points, we developed an aeroponics based maize-AMF co-cultivation system (Figure 5) to study what metabolites secreted in root exudates may play a role during AM symbiosis (Decouard et al. 2025).
 Figure 5: Maize-AMF co-cultivation aeroponic system for studying root exudate and metabolic composition.
Figure 5: Maize-AMF co-cultivation aeroponic system for studying root exudate and metabolic composition. We will especially focus our research on rhizospheric free-living and hyphosphere-associated bacteria. This work will be implemented in three major steps: trapping bacteria and fungi in the rhizosphere (collab J. Fievet, and R. Rincent GQE), creating synthetic communities (Syncom), determining the exudates involved in their recruitment and optimizing the synergistic effect of the SynCom-AMF combination on maize N metabolism. The impact of the bacterial consortia and AMF on maize N metabolism will be characterized.
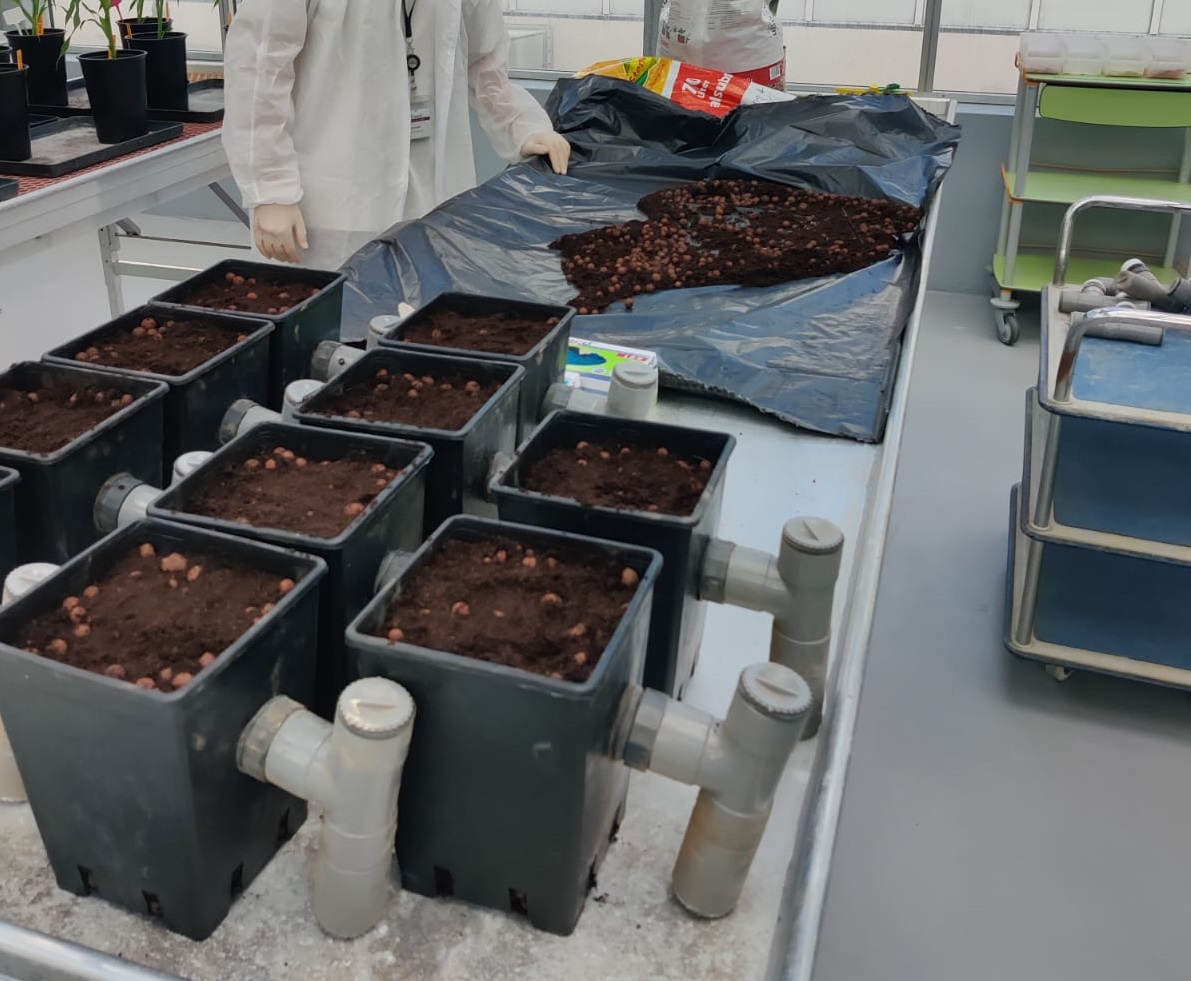

Figure 6: Rhizophere- and hyphoshere-associated bacteria trapping system using maize-AMF association.
C) What are the mechanisms involved in cereal N nutrition under cereal-legume intercropping system in presence of N-fixing legume symbionts and common mycorrhizal networks ? Intercropping systems represent key agroecological strategies to optimize field space and reduce the use of synthetic N fertilizers (Figure 7). We aim to explore the mechanisms by which common mycorrhizal networks redistribute the legume/rhizobium-derived fixed-N towards cereals as well as to explore the impact of such complex interaction on maize primary metabolism.
Figure 7: A quadripartite interaction-system involving maize-AMF-soybean-Bradyrhizobium. .
The cereal crop maize is used in association with the legume crop soybean together with its N-fixing symbiont Bradyrhizobium diazoefficiens and in presence of the arbuscular mycorrhizal fungus Rhizophagus irregularis (Figure 8). Intercropping trials are performed under greenhouse and field conditions (Collab. Carine Palaffre, INRAE Saint Martin de Hinx. The impact of intercropping on the maize and soybean rhizophere microbiome will be investigated.
Figure 8: Maize-soybean intercropping systems in the greenhouse.

Figure 9: Photo of Medicago root with double symbiosis. Staining with ink of Medicago root with nitrogen fixing nodule and arbuscules of the arbuscular mycorrhizal fungus.
Models, tools and methods
The main biological models used in the team are maize, sorghum, soybean and rice for the plant, R. irregularis for the AMF and Bradyrhizobium diazoefficiens as the N fixing symbiont of soybean. Isolated microorganisms are still to be characterized in a collection. Several methodological approaches are used on different organs and at different scales of plant organization and on the microbial partners:
-Physiology, metabolism and molecular genetics of plant N nutrition.
-Microbiology, microbial ecology, culturomics.
-Metabarcoding, metagenomics.
-Phenotyping from cellular to whole plant level under controlled or field conditions.
-Systems biology, multi-omics (transcriptomics, metabolomics, proteomics).
-Metabolic modeling.
Economic and societal issues
The research conducted by the SYNAPS team consists of providing tools and markers to select cultivated plants that use nitrogen more efficiently, thus reducing the excessive use of nitrogen fertilizers that are harmful to the environment. As such, the SYNAPS team is involved in innovative projects involving industrial partners and technical institutes.
Major recent publications (to see more follow this link):
Decouard B, Rigault M, Quilleré I, Boutet S, Adam G, Cueff G, Perreau F, Alunni B, Dellagi B, An aeroponic system to characterize maize root exudates in relation to N and P nutrition and arbuscular mycorrhizal symbiosis, J. Exp. Bot., 76(22) : 6683–6699. PubMed | DOI
Smith NT, Boukherissa A, Antaya K, Howe GW, Mergaert P, Rodríguez de la Vega RC, Shykoff JA, Alunni B, diCenzo GC (2025). Taxonomic distribution of SbmA/BacA and BacA-like antimicrobial peptide transporters suggests independent recruitment and convergent evolution in host-microbe interactions. Microb Genom, 11(4):001380. PubMed | DOI
Urrutia M, Blein-Nicolas M, Fernandez O, Bernillon S, Maucourt M, Deborde C, Balliau T, Rabier D, Benard C, Prigent S, Quilleré I, Jacob D, Gibon Y, Zivy M, Giauffret C, Hirel B, Moing A (2024). Identification of metabolic and protein markers representative of the impact of mild nitrogen deficit on agronomic performance of maize hybrids. Metabolomics, 20(6):128. PubMed | DOI
Chowdhury NB, Simons-Senftle M, Decouard B, Quilleré I, Rigault M, Sajeevan KA, Acharya B, Chowdhury R, Hirel B, Dellagi A, Maranas CD, Saha R (2023). A multi-organ maize metabolic model connects temperature stress with energy production and reducing power generation. iScience, 26(12):108400. PubMed | DOI
Terce Laforgue T, Lothier J, Limami AM, Rouster J, Lea PJ, Hirel B (2023). The Key Role of Glutamate Dehydrogenase 2 (GDH2) in the Control of Kernel Production in Maize (Zea mays L.). Plants, 12(14):2612. PubMed | DOILimami AM, Cukier C, Hirel B (2023). 15N-labelling of Leaves Combined with GC-MS Analysis as a Tool for Monitoring the Dynamics of Nitrogen Incorporation into Amino Acids. Methods Mol Biol, 2642:151-161. PubMed | DOI
Chowdhury NB, Schroeder WL, Sarkar D, Amiour N, Quilleré I, Hirel B, Maranas CD, Saha R (2022). Dissecting the metabolic reprogramming of maize root under nitrogen-deficient stress conditions. J Exp Bot, 73(1):275-291. PubMed | DOIValderrama-Martín JM, Ortigosa F, Ávila C, Cánovas FM, Hirel B, Cantón FR, Cañas RA (2022). A revised view on the evolution of glutamine synthetase isoenzymes in plants. Plant J. PubMed | DOI
Amiour N, Décousset L, Rouster J, Quenard N, Buet C, Dubreuil P, Quilleré I, Brulé L, Cukier C, Dinant S, Sallaud C, Dubois F, Limami AM, Lea PJ, Hirel B (2021). Impacts of environmental conditions, and allelic variation of cytosolic glutamine synthetase on maize hybrid kernel production. Commun Biol, 4(1):1095. PubMed | DOIDellagi A, Quilleré I, Hirel B (2020). Beneficial soil-borne bacteria and fungi: a promising way to improve plant nitrogen acquisition. J Exp Bot, . PubMed | DOI
Dellagi A, Quilleré I, Hirel B (2020). Beneficial soil-borne bacteria and fungi: a promising way to improve plant nitrogen acquisition. J Exp Bot, . PubMed | DOI
Books:
- Philippe Reignault, Ivan Sache, Mathias Choquer, Marie-France Corio-Costet, Alia Dellagi, Frédéric Suffert Phytopathologie, (2023) ISBN-13 : 9782807302884.
- (Chapter) Guellim, A, Hirel, B., Chabrerie, O., Catterou, M., Tetu, T., Dubois, F., Ben Ahmed, H., Kichey, T. (2021) Identification of Physiological Traits Associated with Salinity and Drought Tolerance in Wheat (Triticum durum Desf.).
-(Chapter) Gorawala, P. and Mandhari, S. Eds. Agricultural Research Updates. Vol. 32, Agriculture, Imprints, Nova, Science and Technology Special Topics ISBN: 978-1-53618-974-2.
-(Chapter) Limami, A.M., Cukier, C., Hirel, B., (2023)15N-labelling of Leaves Combined with GC-MS Analysis as a Tool for Monitoring the Dynamics of Nitrogen Incorporation into Amino. In: Couée, I. (eds) Plant Abiotic Stress Signaling. Methods in Molecular Biology, vol 2642. Humana, New York, NY.
Communication visible online :
Invitation by LIED (Laboratoire Interdisciplinaire des Energies de Demain). November 2023 : https://www.youtube.com/watch?v=9gzP28d6CD4 : Title « Microbe mediated plant N nutrition »
Invitation by SNHF-SNHF (Société Nationale d’Horticulture de France). November 2020: https://www.snhf.org/revoir-le-webinaire-3-sante-des-plantes-ressources-naturelles-et-biologie-contemporaine/

Leader:
Alia DELLAGI






